Systematic structure-based analysis of RET variants in MEN2A and Hirschsprung's disease, and the paradoxical co-occurrence of both conditions
Disease Models & Mechanisms, 2026
with Anna Fassler Bakhman, Michal Cohen, and Mickey Kosloff
Interpretable prediction of zinc ion location in proteins with ZincSight
Protein Science, 2025
with Gilad Mechtinger, Gabriel Axel, and Nir Ben-Tal
From binding to catalysis: emergence of a rudimentary enzyme conferring intrinsic antibiotic resistance
MB&E, 2025
with Claudele Lemay-St-Denis, Stella Cellier-Goetghebeur, Maxime St-Aubin, Keigo Ide, Janine N Copp, Soichiro Tsuda, Nir Ben-Tal, and Joelle N Pelletier
A building blocks perspective on protein emergence and evolution
Current Opinions in Structural Biology, 2025
with Yishi Ezerzer, Moran Frenkel-Pinter, and Nir Ben-Tal
A review on this topic
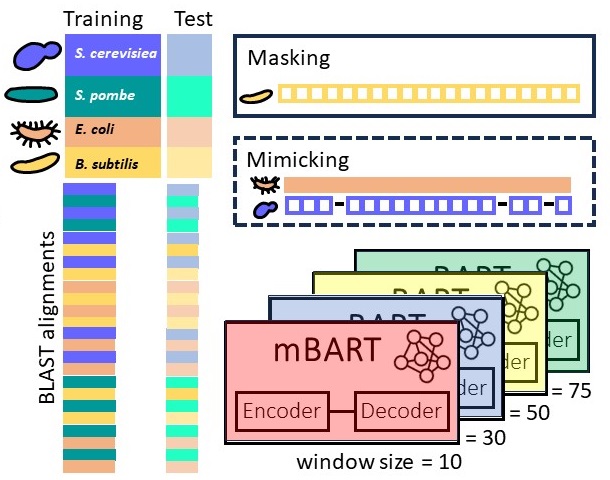
Predicting gene sequences with AI to study codon usage patterns
PNAS, 2024
with Tomer Sidi, Shiri Bahiri-Elitzur, and Tamir Tuller
We study if one can predict codon sequences used by an organism to encode a given amino acid sequence? Codons frequencies vary, a phenomenon known as codon-bias, yet we improve upon frequency-based predictions using contemporary AI tools that learn complex patterns and capture interactions between codons. Because our predictions are tested fairly, on cases not seen during the training process, accurate predictions suggest that these learned patterns are not random, and may be related to the evolutionary process. Our novel AI model for predicting codons is publicly available. That we can better learn codon patterns in high-expression proteins, and in sequences related to housekeeping cellular processes, suggests that the coding sequences of these genes are populated with complex regulatory codes.
PDF | Supplementary data | code | web-server
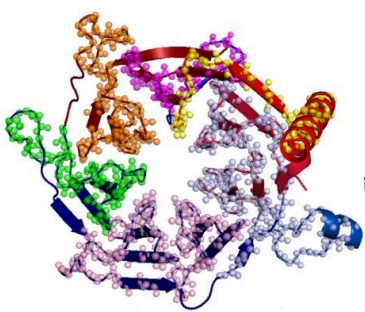
Reused protein segments linked to functional dynamics
MB&E, 2024
with Yigit Kutlu, Gabi Axel, Nir Ben-Tal, and Turkan Haliloglu
Few-shot prediction of the experimental functional measurements for proteins with single point mutations
GEM workshop ICLR, 2024
with Michael Bikman and Rita Osadchy
What can AlphaFold do for antimicrobial amyloids ?
Protein Science, 2023
with Peleg Ragonis-Bachar, Gabi Axel, Shahar Blau, Nir Ben-Tal and Meytal Landau
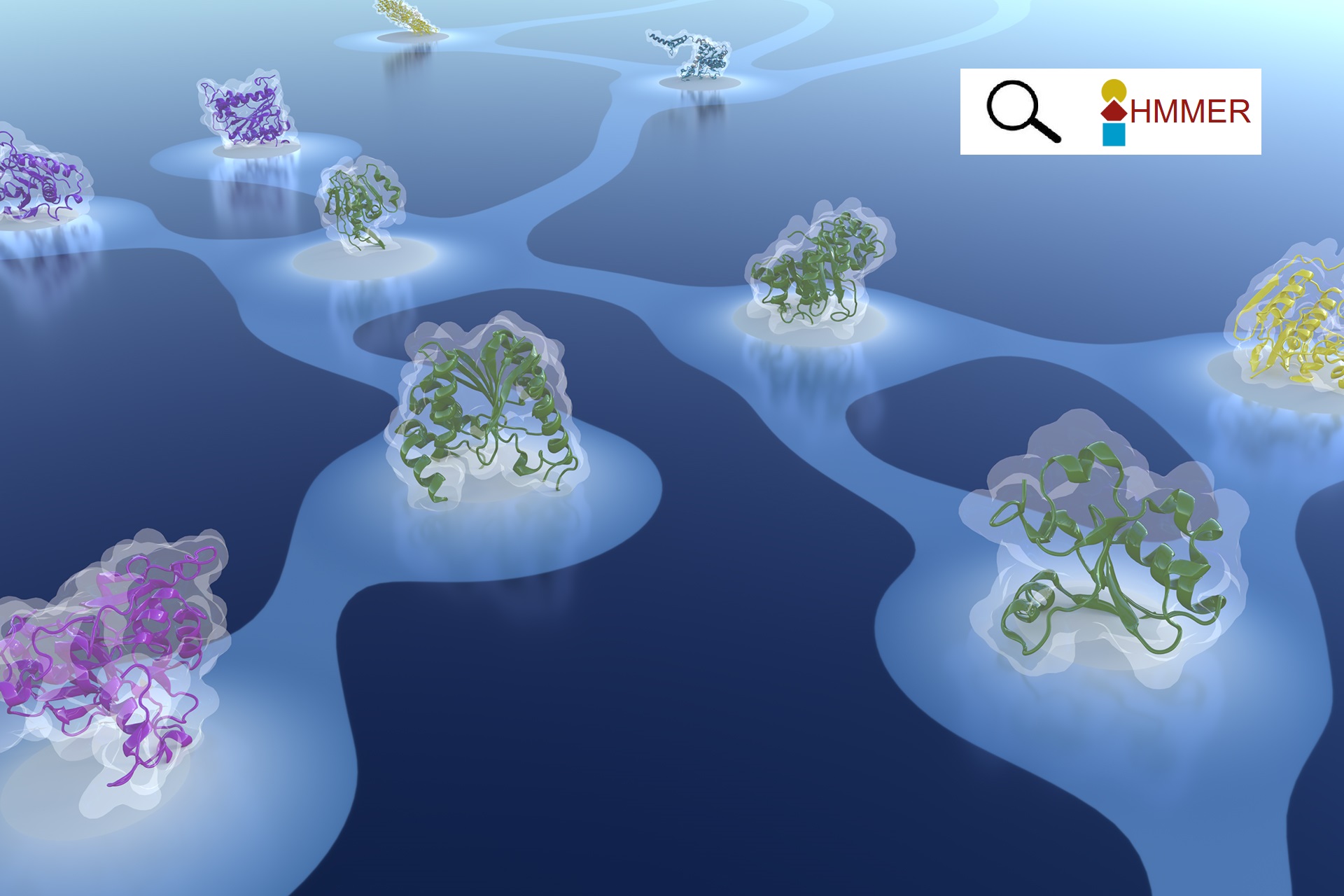

Similar protein segments shared between domains of different evolutionary lineages
Protein Science, 2022
with Kaiyu Qiu and Nir Ben-Tal
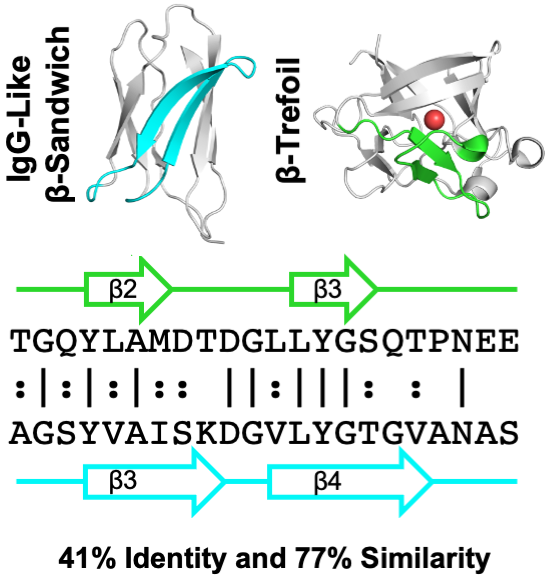
Evidence for the emergence of beta-trefoils by 'peptide budding' from an IgG-like beta-sandwich
PLOS CB, 2022
with Liam Longo and Shawn McGlynn

Adventures on the routes of protein evolution -- in memoriam Dan Salah Tawfik (1955-2021)
JMB, 2022
with Colin Jackson, Agnes Toth-Petroczy, Florian Hollfelder, Monika Fuxreiter, Shina Caroline Lynn Kamerlin, Nobuhiko Tokuriki
Danny Tawfik was my colleague and friend. He is missed.
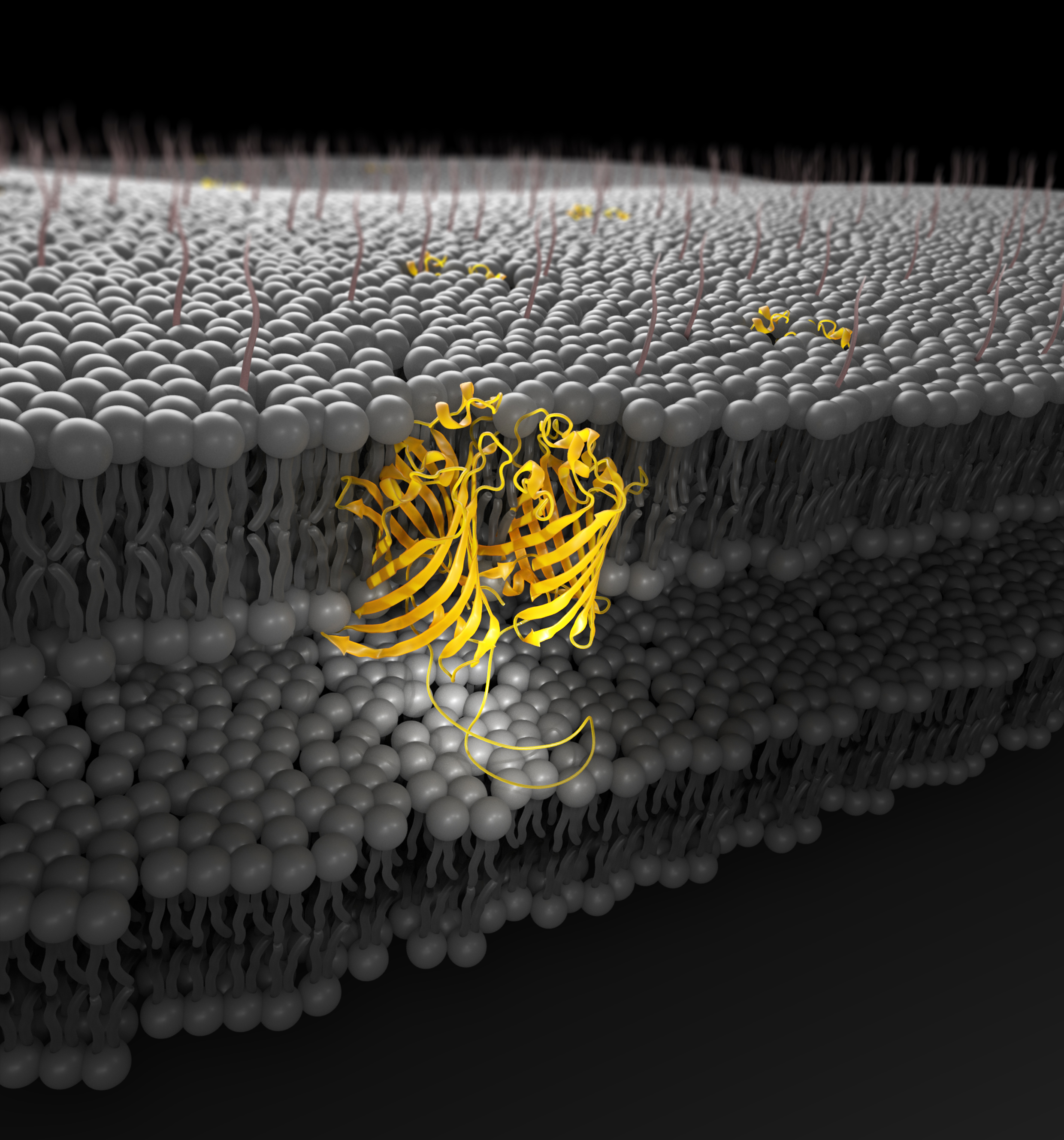

Gram-negative outer-membrane proteins with beta-barrel domains
PNAS, 2021
with Ron Solan, Joana Pereira, Andrei Lupas, Nir Ben-Tal
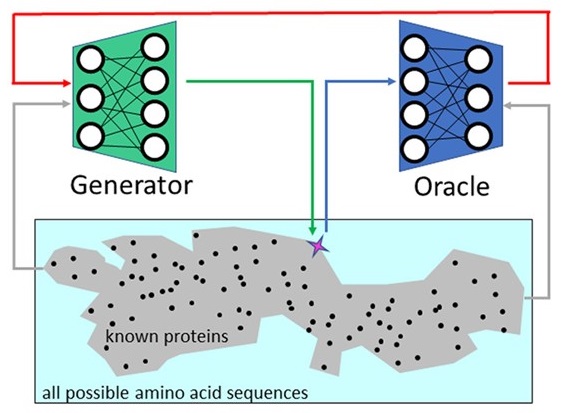
How deep learning tools can help protein engineers find good sequences
J.Phys.ChemB, 2021
with Rita Osadchy
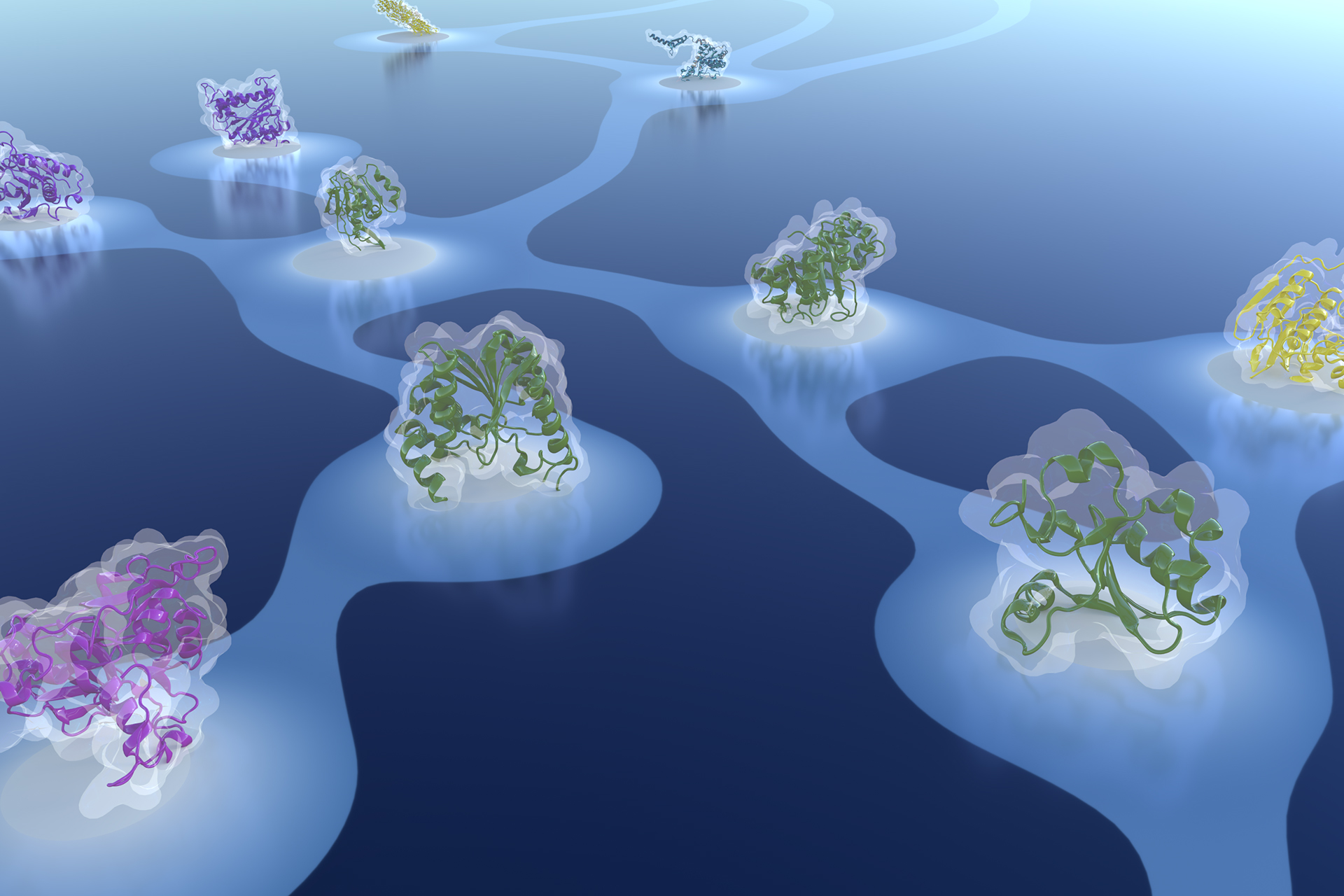
Bridging themes: short protein segments found in different architectures
MB&E, 2021
with Sergey Nepomnyachiy, Dan S Tawfik, Nir Ben-Tal
Searching protein space for ancient sub-domain segments
Curr. Opin. Struct. Bio. , 2021
with nan
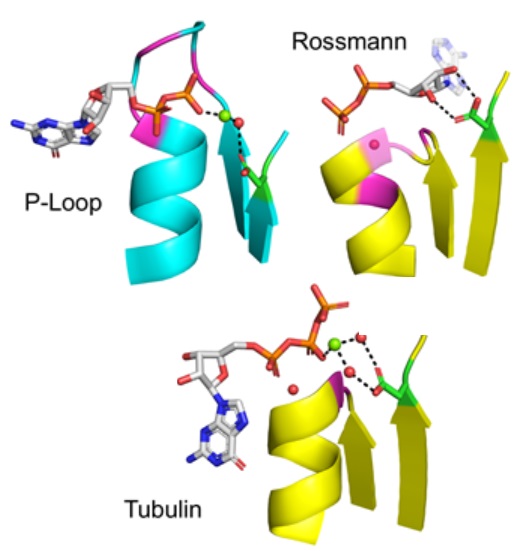
On the emergence of P-Loop NTPase and Rossmann enzymes from a Beta-Alpha-Beta ancestral fragment
eLife, 2020
with Liam M Longo, Jagoda Jablonska, Pratik Vyas, Manil Kanade, Nir Ben-Tal, and Dan S Tawfik
Potential Antigenetic Cross-reactivity Between Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) and Dengue Viruses
Clinical Infectious Diseases, 2020
with Yaniv Lustig, Shlomit Keler, Nir Ben-Tal, Danit Atias-Varon, Ekaterina Shlush, Motti Gerlic, Ariel Munitz, Ram Doolman, Keren Asraf, Liran I Shlush, and Asaf Vivante

Potential In Silico Structural and Biochemical Functional Analysis of a Novel CYP21A2 Pathogenic Variant
International Journal of Molecular Sciences, 2020
with Michal Cohen, Emanuele Pignatti, Monica Dines, Adi Mory, Nina Ekhilevitch, Christa E. Fluck, and Dov Tiosano
On the evolution of protein-adenine binding
PNAS, 2020
with Aya Narunsky, Amit Kessel, Ron Solan, Vikram Alva, and Nir Ben-Tal
To understand how protein-ligand interactions emerged in evolution, we analyzed all protein-adenine complexes of known structure. All of adenine's hydrogen donors and acceptors may facilitate molecular recognition in various binding modes, indicative of convergent evolution. Furthermore, adenine often binds to 'themes', segments of amino acids that are commonly found in proteins, and reported earlier.

Evolutionary pathways of repeat protein topology in bacterial outer membrane proteins
eLife, 2018
with Meghan Franklin, Sergey Nepomnyachyi, Ryan Feehan, Nir Ben-Tal, and Joanna Slusky
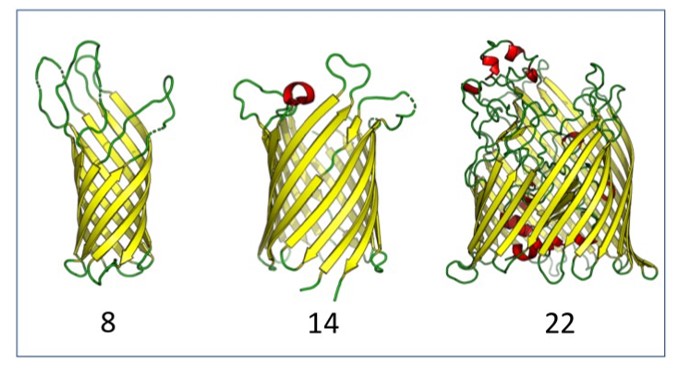
Image from Vikas Nanda's review highlighting this work. Outer membrane proteins (OMPs) are the proteins in the surface of Gram-negative bacteria. These proteins have diverse functions but a single topology: the beta-barrel. Sequence analysis has suggested that this common fold is a beta-hairpin repeat protein, and that amplification of the beta-hairpin has resulted in 8-26-stranded barrels. Using an integrated approach that combines sequence and structural analyses, we find events in which non-amplification diversification also increases barrel strand number. Our network-based analysis reveals strand-number-based evolutionary pathways, including one that progresses from a primordial 8-stranded barrel to 16-strands and further, to 18-strands. We also find that the evolutionary trace is particularly prominent in the C-terminal half of OMPs, implicating this region in the nucleation of OMP folding.
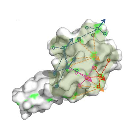
A novel geometry-based approach to infer protein interface similarity
Scientific Reports, 2018
with Inbal Budowski-Tal and Yael Mandel-Gutfreund
We present PatchBag - a geometry based method for efficient comparison of protein surfaces and interfaces. PatchBag is a Bag-Of-Words approach, which represents complex objects as vectors, enabling to search interface similarity efficiently.
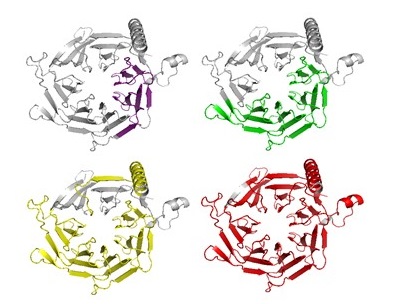
Complex Evolutionary Footprints Revealed in an Analysis of Reused Protein Segments of Diverse Lengths
PNAS, 2017
with Sergey Nepomnyachiy and Nir Ben-Tal
We question a central paradigm: namely, that the protein domain is the "atomic unit" of evolution. In conflict with the current textbook view, our results unequivocally show that duplication of protein segments happen both above and below the domain level among amino acid segments of diverse lengths. Indeed, we show that significant evolutionary information is lost when the protein is approached as a string of domains. Our finer-grained approach reveals a far more complicated picture, where reused segments often intertwine and overlap with each other. Our results are consistent with a recursive model of evolution, in which segments of various lengths, typically smaller than domains, "hop" between environments. The fit segments remain, leaving traces that can still be detected.
Similarity between the Usher Plug and the Repeating Domain of an Ice-adhesin: Evolution via Surface Reshaping
Israel Journal of Chemistry, 2017
with Amit Kessel and Nir Ben-Tal
The PapC usher and MpAFP ice-adhesin feature Ig-like domains, which are similar in shape and sequence but are engaged in vry different functions. We explore how evolution reshaped the surfaces of these two domains to fit to their respective functions.
ConTemplate Suggests Possible Alternative Conformations for a Query Protein of Known Structure
Structure, 2015
with Aya Narunsky, Sergey Nepomnyachiy, Haim Ashkenazy, and Nir Ben-Tal
Protein function involves conformational changes, but often, for a given protein, only some of these conformations are known. The missing conformations could be predicted using the wealth of data in the PDB. Most PDB proteins have multiple structures, and proteins sharing one similar conformation often share others as well. The ConTemplate web server http://bental.tau.ac.il/contemplate) exploits these observations to suggest conformations for a query protein with at least one known conformation (or model thereof). We demonstrate ConTemplate on a ribose-binding protein that undergoes significant conformational changes upon substrate binding. Querying ConTemplate with the ligand-free (or bound) structure of the protein produces the ligand-bound (or free) conformation with a root-mean-square deviation of 1.7A (or 2.2A); the models are derived from conformations of other sugar-binding proteins, sharing approximately 30% sequence identity with the query. The calculation also suggests intermediate conformations and a pathway between the bound and free conformations.
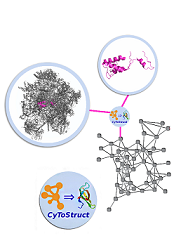
CyToStruct: Augmenting the Network Visualization of Cytoscape with the Power of Molecular Viewers
Structure, 2015
with Sergey Nepomnyachiy, and Nir Ben-Tal
It can be informative to view biological data, e.g., protein-protein interactions within a large complex, in a network representation coupled with three-dimensional structural visualizations of individual molecular entities. CyToStruct, introduced here, provides a transparent interface between the Cytoscape platform for network analysis and molecular viewers, including PyMOL, UCSF Chimera, VMD, and Jmol. CyToStruct launches and passes scripts to molecular viewers from the edges and nodes of the network. We provide demonstrations to analyze interactions among subunits in large protein/RNA/DNA complexes, and similarities among proteins. CyToStruct enriches the network tools of Cytoscape by adding a layer of structural analysis, offering all capabilities implemented in molecular viewers. CyToStruct is available at https:// bitbucket.org/sergeyn/cytostruct/wiki/Home and in the Cytoscape App Store. Given the coordinates of a molecular complex, our web server ( http:// trachel-srv.cs.haifa.ac.il/rachel/ppi/ ) automatically generates all files needed to visualize the complex as a Cytoscape network with CyToStruct bridging to PyMOL, UCSF Chimera, VMD, and Jmol.

Representation of the protein universe using classifications, maps, and networks
Israel Journal of Chemistry, 2014
with Nir Ben-Tal
A meaningful and coherent global picture of the protein universe is needed to better understand protein evolution and the underlying biophysics. We survey the studies that tackled this fundamental challenge, providing a glimpse of the protein space. A global picture represents all known local relationships among proteins, and needs to do so in a comprehensive and accurate manner. Three types of global representations can be used: classifications, maps, and networks. In these, the local relationships are derived, based on the similarity of the proteins sequences, structures, or functions (or a combination of these). Alternatively, the local relationships can be co-occurrences of elements in the protein universe. The representations can be based on different objects: full polypeptide chains, fragments, such as structural domains, or even smaller motifs. Different protein qualities were revealed in each study; many point out the uniqueness of domains of the alpha/beta SCOP (structural classification of proteins) class.
Published in special issue to celebrate Michael Levitt's Nobel prize.

Global view of protein evolution
PNAS, 2014
with Sergey Nepomnyachiy, and Nir Ben-Tal
We've been mentioned in PNAS Highlights (text from there)
Just as the elements in the periodic table can be traced back to the Big Bang, the set of all proteins in terrestrial organisms reflects the history of evolution on Earth. A global view of this so-called protein universe would help reveal how proteins evolve and are related to one another, but empirical evidence exists for relatively few relationships between proteins. Sergey Nepomnyachiy et al. applied network theory to a representative set of all known protein domains drawn from the Structural Classification of Proteins (SCOP) database. The authors represented protein space using two network configurations: a domain network in which edges connect domains the segments of which share similar sequence and structural motifs, and a motif network in which edges connect recurring motifs that lie within the same domains. The authors demonstrate how networks suggest evolutionary paths between domains and provide clues about the mechanisms of protein evolution. The findings offer an approach to representing protein space that could aid protein design, according to the authors.
Redundancy-weighting for better inference of protein structural features
Bioinformatics, 2014
with Chen Yanover, Natalia Vanetik, Michael Levitt, and Chen Keasar
In this study we explore the concept of redundancy-weighted data-sets, originally suggested by Miyazawa and Jernigan. Redundancy-weighted data-sets include all available structures and associate them (or features thereof) with weights that are inversely proportional to the number of their homologs. Here, we provide the first systematic comparison of redundancy-weighted data-sets with non-redundant ones. We test three weighting schemes and show that the distributions of structural features that they produce are smoother (having higher entropy) compared with the distributions inferred from non-redundant data-sets. We further show that these smoothed distributions are both more robust and more correct than their non-redundant counterparts.We suggest that the better distributions, inferred using redundancy-weighting, may improve the accuracy of knowledge-based potentials, and increase the power of protein structure prediction methods. Consequently, they may enhance model-driven molecular biology.
On the Universe of Protein Folds
Ann. Rev. of Biophysics, 2013
with Leonid Pereyaslavets, Abraham O. Samson, and Michael Levitt
In the fifty years since the first atomic structure of a protein was revealed, tens of thousands of additional structures have been solved. Like all objects in biology, proteins structures show common patterns that seem to define family relationships. Classification of proteins structures, which started in the 1970s with about a dozen structures, has continued with increasing enthusiasm, leading to two main fold classifications, SCOP and CATH, as well as many additional databases. Classification is complicated by deciding what constitutes a domain, the fundamental unit of structure. Also difficult is deciding when two given structures are similar. Like all of biology, fold classification is beset by exceptions to all rules. Thus, the perspectives of protein fold space that the fold classifications offer differ from each other. In spite of these ambiguities, fold classifications are useful for prediction of structure and function. Studying the characteristics of fold space can shed light on protein evolution and the physical laws that govern protein behavior.
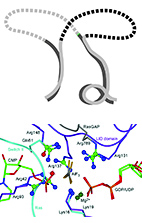
From Protein Structure to Function via Computational Tools and Approaches
Isr. J. Chem. , 2013
with Mickey Kosloff
The 3D structures of proteins are often considered fundamental for understanding their function. Yet, because of the complexity of protein structure, extracting specific functional information from structures can be a considerable challenge. Here, we present selected approaches and tools that were developed in the Kolodny and Kosloff labs to study and connect protein sequence, structure, and function spaces.
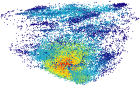
Maps of protein structure space reveal a fundumental relationship between protein structure and function
PNAS, 2011
with Margarita Osadchy
We propose a new method to efficiently create three-dimensional maps of structure space using a very large data set of > 30,000 SCOP domains. In our maps, each domain is represented by a point, and the distance between any two points approximates the structural distance between their corresponding domains. We use these maps to study the spatial distributions of properties of proteins, and in particular those of local vicinities in structure space such as structural density and functional diversity. These maps provide a novel broad view of protein space, and thus reveal new fundamental properties thereof. At the same time, the maps are consistent with previous knowledge (e.g., domains cluster by their SCOP class), and organize in a unified, coherent representation previous observation concerning specific protein folds. To investigate the function-structure relationship, we measure the functional diversity (using the Gene Ontology controlled vocabulary) in local structural vicinities. Our most striking finding is that functional diversity varies considerably across structure space: the space has a highly diverse region, and diversity abates when moving away from it. Interestingly, the domains in this region are mostly alpha/beta structures, which are known to be the most ancient proteins.
A library of protein surface patches discriminates between native structures and decoys generated by structure prediction servers
BMC Structural Biology, 2011
with Roi Gamliel, Klara Kedem, and Chen Keasar
FragBag, a "bag-of-words" representation of protein structure, retrieves structural neighbors from the entire PDB quickly and accurately
PNAS, 2011
with Inbal Budowski-Tal and Yuval Nov
In FragBag, we describe a protein structure by the collection of its overlapping short contiguous backbone segments, and discretize this set using a library of fragments. Then, we represent the protein as a "bag-of-fragments", a vector that counts the number of occurences of each fragment and measure the similarity between two structures by the similarity between their vectors. We use ROC curve analysis to quantify the success of FragBag in identifying neighbor candidate sets in a dataset of over 2,900 structures. The gold standard is the set of neighbors found by six state-of-the-art structural aligners. Our best FragBag library finds more accurate candidate sets than three other filter methods: SGM, PRIDE, and a method by Zotenko et al. More interestingly, FragBag performs on a par with the computationally expensive, yet highly trusted, structural aligners STRUCTAL and CE.
Sequence-Similar, Structure-dissimilar protein pairs in the PDB
Proteins: Structure, Function, and Bioinformatics, 2007
with Mickey Kosloff
It is often assumed that in the Protein Data Bank (PDB), two proteins with similar sequences will also have similar structures. This assumption underlies many Here, we compare sequence-based structural superpositions and geometry-based structural alignments and show that the former provides a better measure of structure dissimilarity. Using sequence-based structural superpositioning we find many examples in the PDB where two proteins that are similar in sequence have structures that differ significantly from one another, usually in direct relation to their function. We conclude that the assumption of two proteins with similar sequences having similar structures is often incorrect and can lead to the loss of structurally and functionally important information.computational studies and structure prediction methods.
VISTAL - A two-dimensional visualization tool for structural alignments
Bioinformatics, 2006
with Barry Honig
VISTAL describes structures as a series of secondary structure elements, and places matched residues one on top of each other colored according to the three-dimensional distance of their Ca atoms.
Using an Alignment of Fragment Strings for Comparing Protein Structures
Bioinformatics, 2006
with Iddo Friedberg, Tim Harder, Einat Sitbon, Zhanwen Li, and Adam Godzik
This work by Iddo and Tim, compares protein structures that are described via strings of fragments from our libraries.
Protein Structure Comparison: Implications for the Nature of 'Fold Space', and Structure and Function Prediction
Curr. Opin. Struct Bio., 2006
with Donald Petrey and Barry Honig
We argue in favor of viewing protein structure space as continuous, with potential structural similarities between any pair of structures. This is different from the traditional perspective in which a structure is in a particular group (denoted fold) and only other structures within that fold are considered as its structural neighbors. We survey recent progress made in the prediction of protein structure and function by relying on these relationships.
Faster Algorithms for Optimal Multiple Sequence Alignment based on Pairwise Comparisons
Lecture Notes in Computer Science (WABI), 2005
with Pankaj K. Agarwal and Yonatan Bilu
We consider the following version of the Multiple Sequence Alignment (MSA) problem: In a preprocessing stage pairwise alignments are found for every pair of sequences. The goal is to find an optimal alignment in which matches are restricted to positions that were matched at the preprocessing stage. We present several techniques for making the dynamic programming algorithm more efficient, while still finding an optimal solution under these restrictions. In our formulation the MSA must conform with pairwise (local) alignments, and in return can be solved more efficiently. We prove that it suffices to find an optimal alignment of sequence segments, rather than single letters, thereby reducing the input size and thus improving the running time.
Comprehensive Evaluation of Protein Structure Alignment: Scoring by Geometric Measures
J. Mol. Biol., 2005
with Patrice Koehl and Michael Levitt
We report a comprehensive comparison of protein structural alignment methods. Specifically, we evaluate six publicly available structure alignment programs: SSAP, STRUCTAL, DALI, LSQMAN, CE and SSM by aligning all 8,581,970 protein structure pairs in a test set of 2,930 sequence diverse protein domains. We follow the traditional path and rely on a gold standard (the CATH classification) and compare the rates of true and false positives using ROC curves. However, due to limitations of this methodology, we also compare the alignments directly, using geometric match measures.
Inverse Kinematics in Biology: The Protein Loop Closure Problem
Int. Jour. Robotics Research., 2005
with Leonidas Guibas, Michael Levitt and Patrice Koehl
We address an inverse kinematics problem in structural biology: the loop closure problem. We describe a procedure for generating the conformations of candidate loops that fit in a gap in a protein structure framework. Our method concatenates small fragments of protein from small libraries of representative fragments. Our approach has the advantages of ab initio methods since we are able to enumerate all candidate loops in the discrete approximation of the conformational space accessible to the loop, as well as the advantages of database search approach since the use of fragments of known protein structures guarantees that the backbone conformations are physically reasonable.
Approximate Protein Structural Alignment in Polynomial Time
PNAS, 2004
with Nathan Linial
Protein structural alignment is a fundamental problem in computational structural biology. Here, we study it as a family of optimization problem and provide a polynomial time algorithm to solve them. We also show an NP-hardness proof of an alternative approach to this problem using internal distance matrices. Lastly, we visualize the scoring function for several pairs of structures.
Protein Decoy Assembly Using Short Fragments Under Geometric Constraints
Biopolymers, 2003
with Michael Levitt
We use the libraries of fragments described below to generate decoys for several proteins. Coupled with a descriminating energy function, decoys are useful for predicting protein structure. It seems that this method works well for all alpha proteins.
Small Libraries of Protein Fragments Model Native Protein Structures Accurately
J. Mol. Biol., 2002
with Patrice Koehl, Leonidas Guibas, and Michael Levitt
We study efficient means of modeling protein structure. Our model concatenates elements from libraries of commonly observed protein backbone fragments into approximate structures. There are no additional degrees of freedom so a string of fragment labels fully defines a three-dimensional structure; the set of all strings defines the set of structures (of a given length). By varying the size of the library and the length of its fragments, we generate structure sets of different resolution. With larger libraries, the approximations are better, but we get good fits to real proteins (less than 1A) with less than 5 states per residue.